Table of Contents
Estimating sequencing depth along contigs
Joran Martijn (August 2025)
After assembling your genome of interest, you may be interested in estimating and/or plotting sequencing depth along the contigs of your assembly. This can be useful for evaluating the quality of your assembly, and spot potential issues. It also just gives you a good idea of the level of your sequencing depth. Maybe its not as much as you thought it was, and you need to sequence more to have more confidence in your assembly.
In any case, at the time of writing, there are two main sequencing technologies used: Illumina (short but highly accurate reads) and Oxford Nanopore / Pacific Biosciences (long but somewhat inaccurate).
It's useful to get an idea of sequencing depth for both these technologies.
Mapping short reads to the assembly
For short Illumina reads, we'll want to use a short read mapper. One of the most used nowadays is BWA (Burrow Wheelers Aligner), developed by Heng Li (who also developed the SAM format, and Minimap, amongst others). Other options are HISAT2 and Bowtie2.
Currently, the best and most computational efficient BWA algorithm is MEM2. It yields identical results to MEM, but is several times faster. To map your short reads to your assembly, you can use the run_bwa_mem2.sh script on Perun.
#!/bin/bash
#$ -S /bin/bash
. /etc/profile
#$ -cwd
#$ -pe threaded 40
#$ -q 256G-batch
# input
base='/scratch3/rogerlab_databases/Fastq_files/Ergobibamus/Illumina/gDNA/qc_seq/trimmomatic-0.39'
R1=${base}/eb_dna_fw_prd.fastq.gz
R2=${base}/eb_dna_rv_prd.fastq.gz
U1=${base}/eb_dna_fw_unp.fastq.gz
U2=${base}/eb_dna_rv_unp.fastq.gz
ASSEMBLY='canu_ergo_contigs.fasta'
# output
PREFIX=dnaseq_vs_canu_medaka_pilon_polish
# load bwa-mem2 software
source activate bwa-mem2
# align dnaseq reads
## first index
bwa-mem2 index -p ${ASSEMBLY%.fasta} $ASSEMBLY
## then align paired reads
bwa-mem2 mem -t 40 ${ASSEMBLY%.fasta} $R1 $R2 | samtools sort --threads 40 -o ${PREFIX}-pe.sort.bam -
## and unpaired reads
bwa-mem2 mem -t 40 ${ASSEMBLY%.fasta} $U1 | samtools sort --threads 40 -o ${PREFIX}-se1.sort.bam -
bwa-mem2 mem -t 40 ${ASSEMBLY%.fasta} $U2 | samtools sort --threads 40 -o ${PREFIX}-se2.sort.bam -
conda deactivate
# indexing bam files
for BAM in *.bam; do
samtools index -@ 40 $BAM $BAM.bai
done
Change the values of the input and output parameters to fit to your assembly and Illumina read files. NOTE that the results of bwa-mem2 are directly piped to samtools sort, meaning that the creation of a SAM file is skipped, and a sorted BAM file is immediately created. This can significantly reduce space on disk.
If you want to merge all three BAM files into a single BAM file, you can make use of samtools cat:
samtools cat ${PREFIX}-pe.sort.bam ${PREFIX}-se1.sort.bam ${PREFIX}-se2.sort.bam > ${PREFIX}-merged.sort.bam
samtools index ${PREFIX}-merged.sort.bam
Important!
Above we didn't pass on any special parameters to bwa-mem2 and samtools. This means it is essentially run in default mode. It turns out that if you are interested in assessing read depth along contigs, this is not exactly how we want to execute things.
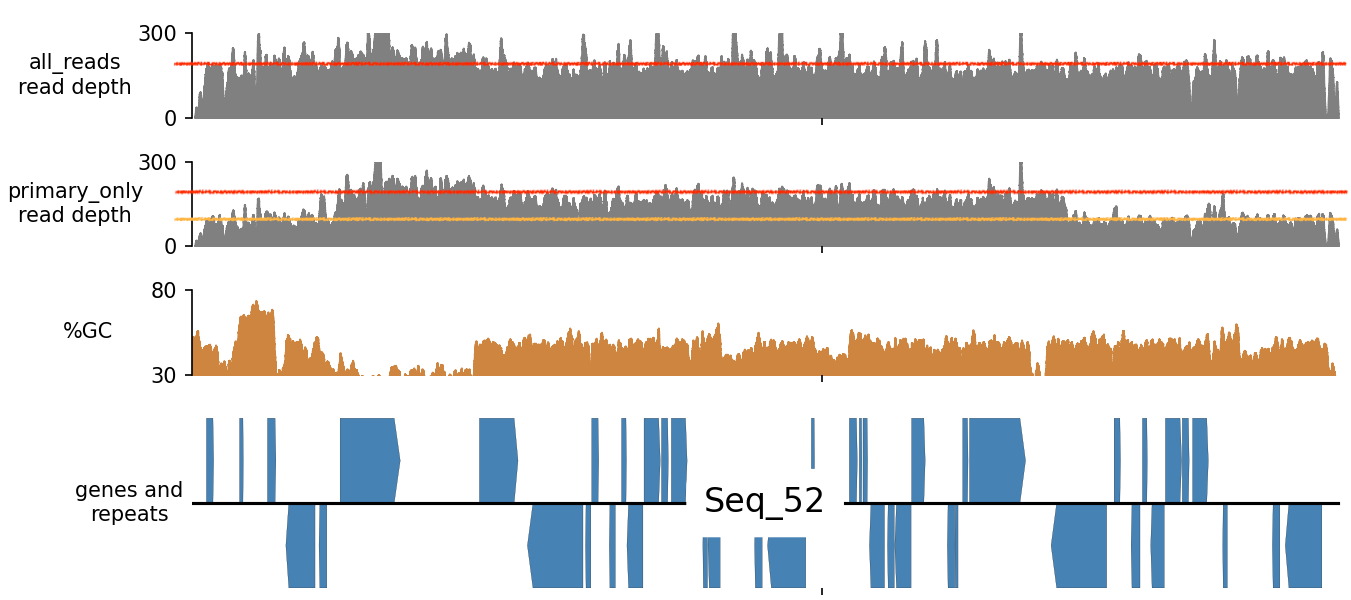
Whenever a sequencing read has sufficient sequence similarity to multiple areas in the assembly, it will report all of these read alignments in the BAM file. The best mapping one will get the label “primary”, whereas the others are considered “secondary”. If you load your BAM file in some software (for example IGV) to visualize sequence depth, both primary AND secondary read mappings will contribute the read depth plot. This gives us a skewed picture of local sequencing depth levels. This is because a single read, which is derived from only one area in a genome, is now contributing to the read depth plot in multiple areas.. See below:
In the picture above, the top coverage or read depth track is drawn using ALL read mappings. Primary and Secondary read mappings. In the coverage track below that, only primary read mappings are used. Suddenly we can see that at the contig edges, read depth is approximately only half what it is compared to the inner part of the contig!
We currently have two hypotheses to explain such odd patterns:
1. There is some biological variation in genome structure within the sampled Blastocystis population. The assembler finds multiple variants of the same homologous region in the read data, and represents this area multiple times in different parts of the assembly. Here, the for example 3' edge region of contig Seq_52 is also found on some other contig (not shown).
Short reads will equally match both areas, and thus their coverage is split between those two, while short reads from unique areas only have one clear mapping location. As a result, the edge regions that occur twice get half the short read coverage relative to the inner regions that only occur once in the assembly!
2. There is some artificial variation in the assembly, because some long reads were so enriched in errors that the assembler didn't recognize they'd overlap with other long reads that stem from the same chromosome region. And these errors somehow persisted through the polishing process.
In any case, now that we've filtered out the secondary mappings, we can at least see that something fishy is going on here! If we didn't, we'd falsely get the impression that everything is fine and dandy.
Thus, we need to prevent secondary mappings to be reported in the BAM file. This can be achieved by editing our bwa-mem2 / samtools sort command by adding the samtools view -b -F 256 in the piped command:
## then align paired reads
bwa-mem2 mem -t 40 ${ASSEMBLY%.fasta} $R1 $R2 | samtools view -b -H -F 256 | samtools sort --threads 40 -o ${PREFIX}-pe.sort.bam -
The -F 256 flag means that any read alignment detected by bwa-mem2 that has the bitwise flag 256 set will be omitted in the final BAM file output. The 256 bitwise flag is code for the read alignment being secondary.
Alternatively, if you already have a BAM file, you can either apply the same command on that BAM file:
samtools view -b -H -F 256 <BAM> > <NEWBAM>
Or you can use bamtools :
bamtools filter -in <BAM> -isPrimaryAlignment true -out <NEWBAM>
Plotting sequencing depth along the assembly
There are multiple ways to do this, but perhaps the most straightforward way is to use the Integrated Genomics Viewer (IGV).
First load your genome with Genomes→Load Genome from File…, and then load the BAM file with File→Load from File…